Note: post updated, see below!
This post is a bit all over the place, but bear with me. I've had a very trying day trying to combine paired end reads into longer single end reads. While currently not having the most success myself, I gained a few insights along the way that might help out some other people a little, if only to see how someone did it badly.
This post is a bit all over the place, but bear with me. I've had a very trying day trying to combine paired end reads into longer single end reads. While currently not having the most success myself, I gained a few insights along the way that might help out some other people a little, if only to see how someone did it badly.
Let's set the scene; I recently ran a small sample of four pooled amplicons on the same index on a paired-end, 2x251bp MiSeq run (I luckily managed to squeeze into a friend's run, as I just wanted to test the protocol was working).
These amplicons should be roughly different lengths: two amplicons should be around 430 bp, while the other two will be a bit shorter, around 320 bp, so they should have overlaps around 70 and 180 bp long respectively.
Stitching these overlapped reads would make a lot of my downstream processing easier, and seems like it should be fairly easy, right? There's loads of programs and scripts designed to do just that (summed up nicely in this nice post on The Genome Factory).
If only life were that easy.
I first tried what seemed to be the big three: FLASH, COPEread and PANDASeq. Getting the things up and running proved to be no mean feat - the classic of games of find-the-dependency, excavating old binaries and correcting the typos in the installation commands of course only fuelling my analytical mindset.
I tried COPEread first. It's a bit tricker than it first might seem, as in order to work fully you first need to generate k-mer frequency tables from the files you wish to overlap, using the bundled program kmerfreq on a text file containing the names of the fastq files, like so:
kmerfreq -k 16 -t 3 -q 33 -p outputfile filenames.lst
Here -k is the length of the kmers, -t the number of threads to use, and -q 33 indicates that I'm using phred scores starting from ASCII value 33, hence (newer) Illumina values - COPE really seem to like making the old value of 64 as the default. Also note that I had to use kmers of 16, below the default of 17, as 17 or above is seemingly beyond the abilities of the memory of my 8 GB machine.
Running COPEread on its most basic settings looks like this.
cope -a R1.fq -b R2.fq -o merged.fq -2 leftovers_R1.fq -3 leftovers_R2.fq -s 33 -m 0
(Note that it seems a bit particular about certain parameters being filled, and might not give a sensible error message)
However, this manages to join less than a percent for my amplicons. Let's try beefing it up to the full mode, using the kmer frequency table made above:
cope -a R1.fq -b R2.fq -o merged.fq -2 leftovers_R1.fq -3 leftovers_R2.fq -s 33 -m 3 -k 16 -t freqtable.freq.cz -f freqtable.freq.cz.len
Still we're getting effectively no reads joining. Maybe COPEread isn't the one.
Let's try PANDAseq. Let's please try PANDAseq. Why oh why did this installation take me so long?
I tried so many ways to get this working, that I can now no longer seem to even run it properly, as there are libraries and binaries strewn across directories all over my drive. However, I ran up against the seemingly well known problem that PANDAseq is a fussy eater - https://github.com/neufeld/pandaseq/issues/7, meaning it wouldn't accept my data, even using the -B flag.
No problem, I can just dress my IDs up with fake barcodes comme ça, before running PANDAseq on them:
sed -i '1~4s/$/\ 1:N:0:TAGACA/' R1.fq
sed -i '1~4s/$/\ 2:N:0:TAGACA/' R2.fq
pandaseq -f R1.fq -r R2.fq -u unpaired.fq
Now it seems to recognise the data as valid, but doesn't do anything afterwards. It just finds no pairs, and outputs nothing into the unpaired file. According to the bundled pandaseq-checkid, my IDs are still 'BAD', but as far as I can tell they shouldn't be. However, by this point I'd spent far too long plugging away at just one program when I still had more potentials, so maybe the next wouldn't be so pernickity.
(In fairness to the PANDAseq devs, they do seem to be interacting with the users and fixing the bugs - I'm sure if I chased this I could get some help).
On to FLASH, which seems to be a bit of a favourite over on SEQanswers. Nice and simple default options:
flash R1.fq R2.fq
This combined 30% of the reads! Admittedly not really as high as we'd want, but certainly the best so far.
I spent a long time flag-fiddling with FLASH, with no real improvement. Here's my most redundantly parameterised command, stipulating the minimum/maximum overlap sizes, length of reads and amplicons, with standard deviation:
flash -m 30 -M 220 -r 502 -f 376 -s 56 s1.fq s2.fq
Still wobbling around the 30% mark. Getting pretty frustrated by now, so I asked the good people over at SEQanswers, and received the great suggestion to demultiplex my sample by amplicons.
Knocked up a quick python script to scroll through the R2 fastq (which contains the primers that differentiate the amplicons) and bin reads into different files based on which sequence it finds (in a post to follow!). Once I've split the R2 into four, I can harvest the ID line of each with sed, remove the @ and then use fastqselect.tcl to pull the appropriate matched reads out of R1, e.g.:
sed '2~4d;3~4d;4~4d' R2.fq > idR2
...
for i in id*; do echo $i; sed -i 's/@//g' $i; done
Now I get something really interesting. The two amplicons in my pool that were longer (~430 bp, ~70 bp overlaps) combine great, over 90% assembling. The shorter amplicons (~320 bp, ~180 bp overlaps) however resolutely refuse to combine at all, even though if I pick a few mate reads and BLAST them against each other I can see ~200 bp overlaps with 99-100% identity.
I don't know why I couldn't get COPEread to work, I don't know what ID format PANDAseq was looking for, and I don't know why FLASH wont combine rather extreme long-ish reads where the overlap makes up a rather large percentage of each read.
Sadly my time is tight; I don't anticipate this to be a regular feature of my pipeline (as my usual amplicons are too long to overlap) and I only needed a couple of joined files to test the analysis I wanted to try out, so I'm yet to pursue this. Truthfully I probably spent more time to do this than I should have, because I'm stubborn and it seems like something that should be simple.
Because it should be simple, right?
If you can see something obvious I'm doing wrong, please let me know in the comments.
Post-script:
I did also try a few other methods.
Pear said that the number of reads in each file did not match (when clearly they did).
mergePairs.py looked like it was working, but inspection of the merged files it was producing shows a number of clearly junk reads stuck together. Also obviously slower than the C based approaches above.
SeqPrep never installed, but as it trims and assembles in one I'm not sure if it's appropriate for me (as I have random sequence at both ends of the amplicons).
Bearing in mind that all of these presumably work for at least some (if not most) people, I'm aware that the fault probably lies with my data, which I expect is longer and more overlapped than most of these programs were designed for. That said, if I can align some of these myself just by eye, I don't see why the leading software in the field can't.
UPDATE (19th July 2013)
After throwing this post up on Twitter, I got a bit of feedback from my local sequencing guru Tony Brooks and doyen of bioinformatics Nick Loman, which prompted me to have a second crack at my difficult overlaps.
First off, a correction: earlier in this post I mentioned that I'd tried FLASH using both the -M and -rfs flags all at once. I had tried them individually (either -M or -r -f -s), with equal success.
The main thing to come out of the discussion was the obvious, which is that I don't really need to combine the reads of the shorter length amplicons, as the read length already spans the entire insert (fig. 1).
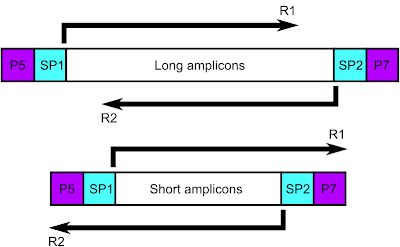 |
| Figure 1: Schematic of short versus long amplicons |
However, I'd still like the possibility to combine these reads. For one thing, it'd be nice to have just one pipeline which I can roll out across all my amplicons, without my having to demultiplex them as I did here. More importantly, the quality of the reads starts to dip the further through the read we get, so combinining the reads would allow me to use the higher confidence base calls from the other direction.
The major advice given was that I should trim the adaptors off my reads, which should help with the combinining. Out comes Trimmomatic, an excellent bit of kit which seems to be a bit of a trimming-favourite.
Frustratingly, the success I get from Trimmomatic mirrors that which I see with FLASH:
java -jar ~/PATH_TO_FOLDER/trimmomatic-0.30.jar PE -phred33 -trimlog trimlog R1.fq R2.fq trimpairR1.fq trimUNpairR1.fq trimpairR2.fq trimUNpairR2.fq ILLUMINACLIP:TruSeq3-PE.fa:2:30:10 LEADING:3 TRAILING:3 SLIDINGWINDOW:4:20 MINLEN:80
When applied to all four of my demultiplexed samples, almost all of the reads in R2 fail to pass muster for two of the amplicons - any guesses which? The shorter amplicons not only refuse to merge, they refuse to trim to assist merging! I initially put this down to a slightly worse quality profile in the R2 reads, but the difference between the quality of R1 and R2 is about the same among both longer and shorter amplicons.
It was around this time that Eric Biggers, one of the FLASH software developers, replied to an email I'd sent him earlier detailing my problem, with a couple example reads which I thought should align. Here's his reply:
"The problem is that FLASH is looking for overlaps where read 2 ends at some position in read 1, whereas in the two example pairs you gave, read 2 ends before read 1 even begins.So your data looks something like:read_1 --------------------->
read_2 <--------------------which FLASH doesn't expect, since those are, in effect, outward-facing pairs rather than the inward-facing pairs it expects. In both cases the sequence at the end of read 2 (at the head of the arrow) is similar, so I assume it's an adapter sequence. In short, FLASH wasn't originally designed to handle cases like this, but after seeing this I'm considering modifying it to look for such overlaps, especially since I recall someone else having a similar problem."
There you have it; it makes so much sense, I feel silly for not having thought of it sooner (hey, I'd had a long day) - it can't find the overlap, because the sequence at the end of the read doesn't appear in the other read!
Combined with the trimming advice I'd received, there was then a simple way to rectify this problem; just reduce the read length to make my data into inward-facing pairs. The FASTX-Toolkit is great for this kind of thing (just remember to use the -Q33 flag to make sure it's looking for the right phred scale - I still can't fathom why that doesn't feature in their manual!).
fastx_trimmer -Q33 -f 1 -l 175 -i R1.fq -o shortR1.fq
fastx_trimmer -Q33 -f 1 -l 175 -i R2.fq -o shortR2.fq
flash shortR1.fq shortR2.fq
Eureka! I now get as good merging of pairs for the shorter amplicons as I saw with the longer! So, were I to roll this out to my un-demultiplexed amplicon pool data I imagine I would run FLASH on the original files, then trim any reads that don't merge before re-running FLASH on those.
Big thanks to Eric Biggers, Tony Brooks and Nick Loman and all the people on SEQanswers for their help.
Big thanks to Eric Biggers, Tony Brooks and Nick Loman and all the people on SEQanswers for their help.